Theoretical and Experimental Investigations of the Basal Ganglia
How does the lack of dopamine produce the symptoms of Parkinson's Disease?
How does excessive dopamine in response to drugs of abuse produce addiction?
We created a theta burst stimulation paradigm for inducing LTP in normal magnesium. We demonstrated changes in the ability to induce striatal synaptic plasticity in rats that have learned to navigate a T-maze, and found changes correlated with training stage.
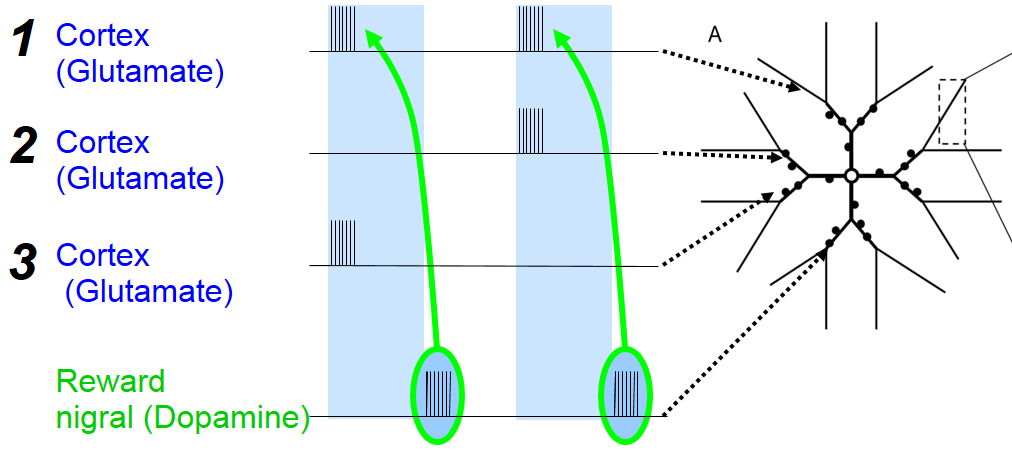
We demonstrated that calcium dynamics predicts the change in synaptic weight for three different STDP protocols.
Large scale simulations of striatal network models reveals that the abnormal connectivity between striatal neurons causes beta oscillations. Dopamine depletion weakens the lateral inhibitory connections, and strengthens the feedforward inhibition from fast spiking interneurons. Synchronizing input to the gap junction connected FSIs produces beta band oscillations. A prediction of the model is that blocking gap junctions will normalize striatal activity and restore normal movement in Parkinson's Disease. Read more about this research here.
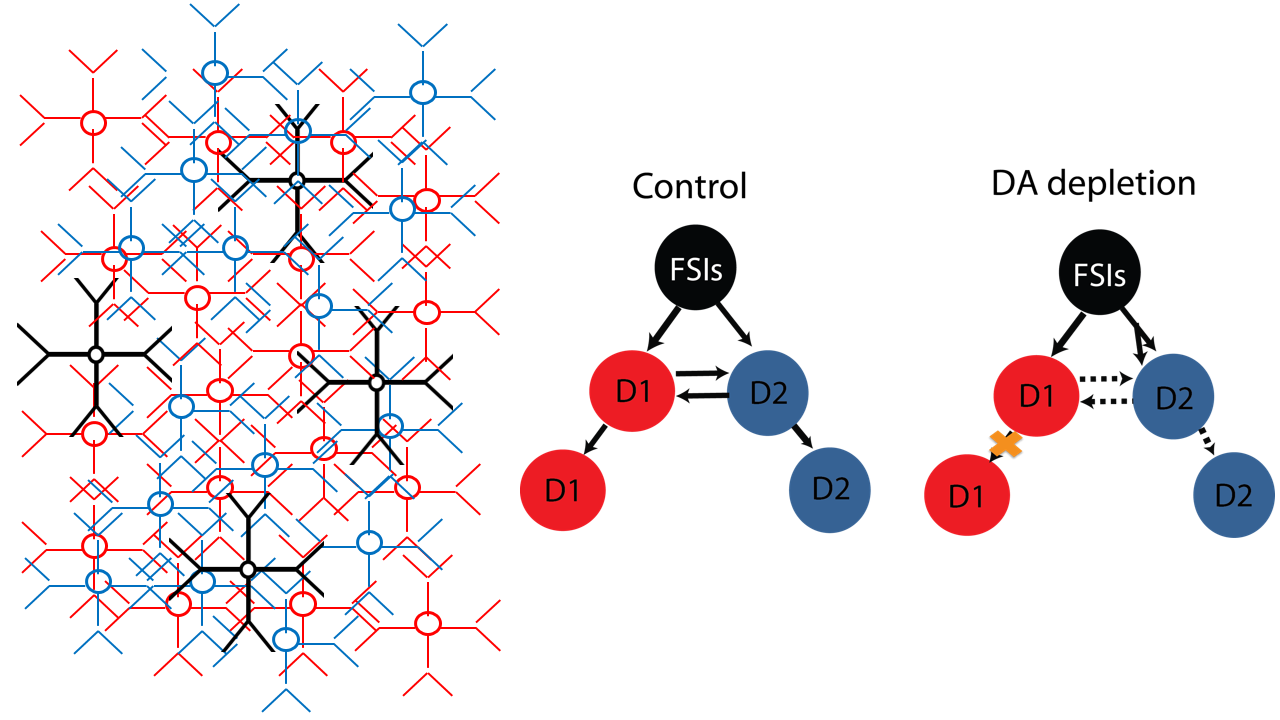
Striatal Network Model
Mechanisms underlying Hippocampal LTP
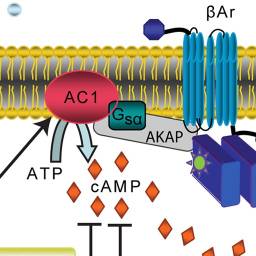
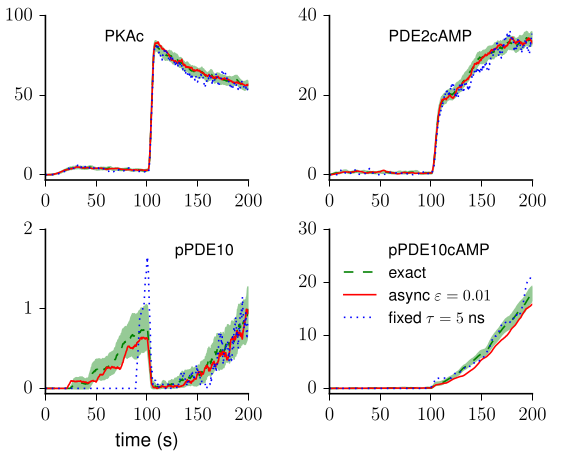
 Our favorite neural simulator:
Our favorite neural simulator: 